Guest Blogger Prof Sally Bloomfield (bio below) writes…In proposing the hygiene hypothesis in 1989, Dr David Strachan suggested that lower incidence of early childhood infections could explain the 20th century rise in allergic diseases. This was based on data showing that larger family size appears to protect against hay fever. Strachan suggested that smaller families provided insufficient infection exposure because of less person to person spread of infections – but also because of “improved household amenities and higher standards of personal cleanliness”. From this the notion that “we have become too clean for our own good” has arisen.
Most experts now agree that the “hygiene hypothesis” is a misnomer. Although the basic concept is still seen as correct, the link to infectious disease and hygiene is now largely discounted. A number of refinements to the hypothesis offer a more plausible explanation. The Old Friends (OF) Mechanism was proposed by Graham Rook in 2003. He proposed that the required exposures are not childhood diseases such as colds, flu, measles, norovirus, which have evolved only over the last 10,000 years, but the microbes we co-evolved with in hunter-gatherer times when the human immune system was developing. These Old Friends include largely non-harmful organisms such as helminths, commensal microbiota and environmental saprophytes.
Although these microbes are still there, through lifestyle changes we have gradually lost our exposure to them. Improved water, sanitation and food quality, although protective against infections, have also inadvertently reduced exposure to Old Friends which occupy the same habitats. The decline in natural childbirth in favour of caesarean section, and bottle instead of breast feeding is another likely factor. Reduced exposure to our outdoor environment has also occurred – we now spend up to 80% of our time indoors. Also, antibiotics may alter our interaction with microbes leading to reduced diversity of human gut microbiota.
Despite a shift in scientific thinking, the so-called “hygiene hypothesis” is still widely accepted in the public domain and still discussed in terms of concepts such as ‘eat dirt’, ‘too clean is bad’, or ‘sterile homes’. The lay audience, and even the U.S. FDA attribute the problem to ‘the extremely clean household environments often found in the developed world’.” (See headlines below!)
What is overlooked is the fact that the relationship between household or personal cleanliness and development of allergies has never been properly investigated. At last – just this week, we have seen publication of the first study to directly evaluate this issue. From the study, Erika von Mutius, a highly respected researcher in this field, concludes – No. “Development of allergies and asthma was not related to cleaning activities”.
Methods: The study involved a birth cohort of 399 participant families recruited in urban and suburban regions of Munich, Germany, between Oct 1999 and Dec 2001. A telephone questionnaire interview comprising 31 questions was carried out to assess cleaning habits and cleaning frequencies in the homes, the use of detergents and the personal cleanliness of the child. In addition, 13 lifestyle factors and home characteristics were obtained from a self-administered questionnaire. Questions about the child’s health focused on respiratory and allergic problems. Bacterial markers of home cleanliness were assessed in samples of floor and mattress dust.
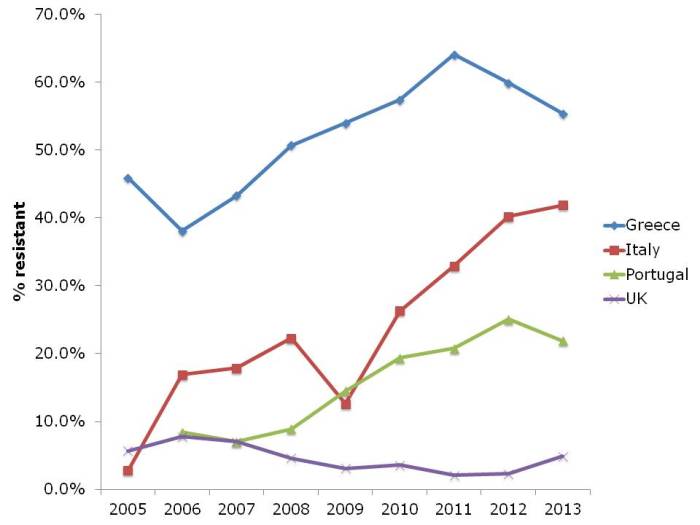
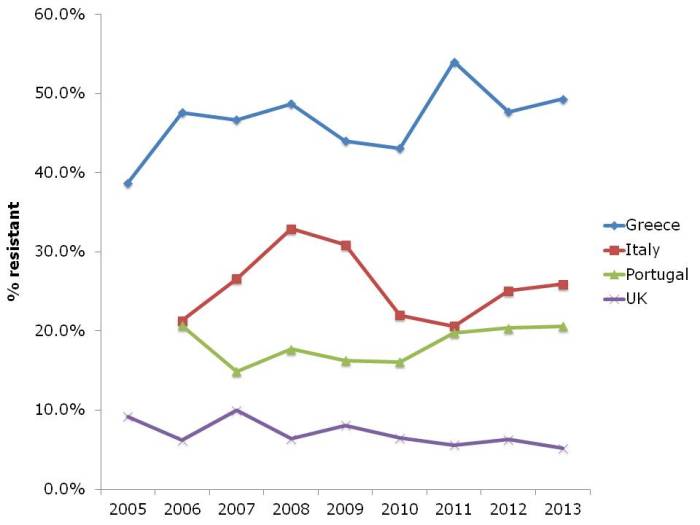
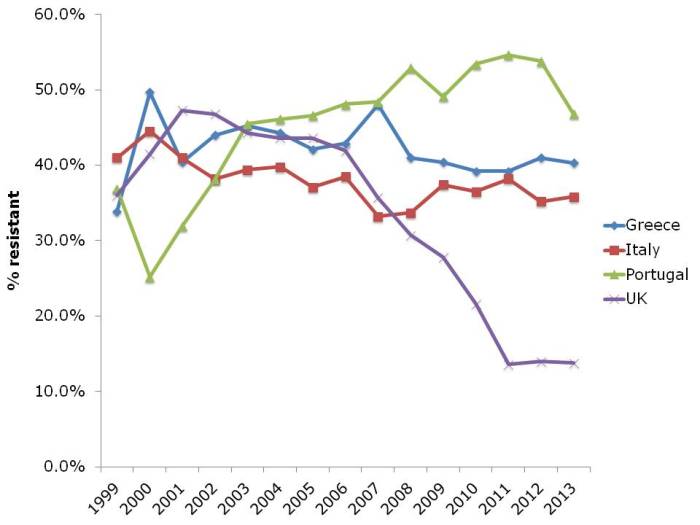
Results and conclusions: As found by other workers, bacterial exposure in house dust was found to be associated with reduced risk of childhood allergies. In turn, personal cleanliness, such as washing hands, and home cleanliness were objectively reflected by dust parameters. However, neither personal nor home cleanliness were associated with protection from asthma and allergies (see flow chart, below).
Note: In this study, cleanliness was substantiated by rather unspecific dust measurements. The findings however suggest that allergy protection operates through as yet unknown exposures, not assessed by unspecific markers. Future studies will require more detailed microbial analysis. Whether these microbes are affected by cleaning remains to be elucidated, but findings with unspecific markers suggest that normal cleaning does not affect permanent microbial colonization of indoor environments.
If, as now seems, allergies are not the price we have to pay for protection from infection, two fundamental questions need to be addressed:
- “How can we develop an approach to hygiene, which helps to reconnect us with the necessary microbial exposures, whilst also protecting us against infectious diseases? “
- How do we change public understanding about the difference between “cleanliness” (absence of visible dirt) and “hygiene” (protecting against infectious diseases)?
The answer to the first question is a return to basics, which means promoting “targeted hygiene”. This means identifying the critical points in the chain of infection transmission and applying effective hygiene procedures at the appropriate times to prevent further spread. Appropriate times are those associated with activities such as food and water hygiene, respiratory hygiene, toilet hygiene, laundry hygiene and so on.
Dispelling the misconceptions is a real challenge, made more difficult by the fact that people tend to think that hygiene and cleanliness is the same thing (i.e “if it looks clean it must be germ free”). It is possible that this is best done by promoting a more constructive approach i.e . stressing that getting dirty is healthy, but hygiene is vital in the times and places that matter.
Further Reading: Bloomfield SF, Stanwell-Smith R, Rook GA. 2013. The hygiene hypothesis and its implications for home hygiene, lifestyle and public health: summary.
The study can be found at: Am J Respir Crit Care Med. 2015 Jan 13. [Epub ahead of print] Asthma and the Hygiene Hypothesis – Does Cleanliness Matter? Weber J, Illi S, Nowak D, Schierl R, Holst O, von Mutius E, Ege MJ.
Guest blogger bio:
Dr Sally Bloomfield is an Honorary Professor at the London School of Hygiene and Tropical Medicine. She is also is the Chairman and Member of the Scientific Advisory Board of the International Scientific Forum on Home Hygiene (IFH). Through these roles Professor Bloomfield continues to develop her work in raising awareness of the importance of home hygiene in preventing the transmission of infectious disease, and developing and promoting home hygiene practice based on sound scientific principles. She is also working to develop understanding of “hygiene issues” such as the “hygiene hypothesis” and “antimicrobial resistance”.
Professor Bloomfield’s background is in healthcare and infectious disease. She has a degree in Pharmacy, and PhD in Pharmaceutical Microbiology from the University of Nottingham. Sally was previously a Senior Lecturer in Pharmaceutical Microbiology at Kings College London (1995 – 1997) and a Hygiene Liaison manager at Unilever Research Port Sunlight UK (1997 – 2001). She has published 100 research and review papers on the subject of home hygiene and the action and mode of action role of antimicrobial agents.