We know that many vegetive bacteria can survive on dry surfaces for longer than you might think. For some Enterococcus species, this capability is nothing short of extraordinary. In one study, Enterococcus dried onto a surface was still viable 4 years (yes FOUR YEARS) later! How can this be? No nutrients, no water (other than ambient humidity), and not an endospore former. A recent paper in the JHI may have some answers: Enterococcus is able to form dry surface biofilms and these contain viable Enterococcus many months after inoculation, regardless of species and substrate.
Continue readingenvironmental contamination
IPS Journal Club: Inhaler vs. nebuliser and the risk of SARS-CoV-2 dispersal
I’ve written this post in preparation for Wednesday’s IPS Journal Club (register here). The paper that I have chosen for the Journal Club is this one in the Journal of Infection Prevention, comparing the risk of SARS-CoV-2 dispersal through the air with an inhaler vs. nebuliser.
Continue readingCandida auris and surface survival
Candida auris is an emerging threat to healthcare facilities worldwide. Recent, worrying, data from the US suggests that prevalence is increasing rapidly. So, we need to make sure we have every prevention base covered to reduce the chances of cross-transmission. C. auris seems to be quite an environmental organism – and a recent JHI study confirms this, showing extended survival on surfaces and tolerance to low concentrations of some biocides.
Continue readingDeveloping antimicrobial “smart surfaces” to tackle HCAI and AMR
I participated in a launch event by the Institute of Molecular Science and Engineering (IMSE) at Imperial College London yesterday for a new white paper on developing “smart surfaces” to tackle HCAI and AMR.
Dispersal of CPE from contaminated sinks and drains: a refection from Infection Prevention 2019
I’ve spent the last couple of days up in Liverpool for Infection Prevention 2019. One of the highlights was a talk by Dr Paz Aranega-Bou on the issues around contamination of sinks and drains. Paz flagged a paper just published in JHI investigating the dispersal of CPE in a sink/drain test risk at PHE, showing the CPE can make its way from contaminated drains to sink and surrounding surfaces via splashback.
CPE in drains: a light at the end of the drain pipe?
We have been posting for a while about the emerging recognition of CPE contamination of drains in clinical settings, which seems to be fueling some CPE transmission. Until now, there’s been plenty of publications identifying the problem, but very few presenting a solution. In fact, attempts to tackle CPE contamination of drains have had moderate impact, at best. A new short study in ICHE illustrates the potential of a foaming hydrogen-peroxide based disinfectant to tackle contamination with resistant Gram-negative bacteria in drains.
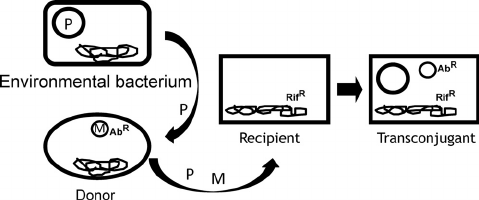
Mcr-1 plasmid-mediated colistin resistance genes in environmental Enterobacteriaceae
An interesting new Italian study has identified the mcr-1 gene, a plasmid-mediated colistin resistance gene, in 8% of environmental Enterobacteriaceae isolates. This suggests that environmental Enterobacteriaceae and perhaps even environmental surfaces themselves could be important reservoirs in the spread of mcr-1 and colistin resistance.
CPE contamination of hospital wastewater: smoking gun or innocent bystander?
A recent US study has investigated CPE contamination of sinks, drains, and wastewater. Carbapenemase-producing bacteria were identified throughout the drainage and water system, from drains in patient rooms, right through to wastewater sampled through manholes adjacent to the hospital. My main question in all of this is whether this huge reservoir of carbapenemases in hospital wastewater is a risk for patients. The lack of genetic similarity between isolates in hospital wastewater and isolates from patients suggest not, but I suspect there’s an indirect link and these carbapenemases find their way into isolates affecting humans, which is supported by genetic links between the plasmids carrying the carbapenemases.
HIS Spring Meeting: ‘Contaminated surfaces: the missing link’
Thought I’d share some key points from the 2016 HIS Spring Meeting.
Outlining the problem(s)
Prof Gary French kicked off the meeting with a (sic) historical perspective, describing how the perceived importance of the environment in transmission has oscillated from important (in the 40s and 40s) to unimportant in the 70s and 80s to important again in the 2000s. Gary cited a report from the American Hospital Association Committee on Infections Within Hospitals from 1974 to prove the point: ‘The occurrence of nosocomial infection has not been related to levels of microbial contamination of air, surfaces and fomites … meaningful standards for permissible levels of such contamination do not exist.’ Gary covered compelling data that contaminated environmental surfaces make an important contribution to the transmission of Gram-positive bacteria and spores, highlighting that C. difficile in particular is a tricky customer, not helped by the fact that many ‘sporicides’ are not sporicidal!
Up-date on M. chimaera
More and more reports and guidance (Ref) appear with regard to Mycobacterial infections associated with heater cooler units used during thoracic surgery. As mentioned in this blog before, the infections are attributed to aerosol generated by the contaminated heater cooler units that are located in or adjacent to the operating room (Ref).
Just now, researchers published 10 patients with disseminated Mycobacterium chimaera infections subsequent to open-heart surgery at three (CH, GER, NL) European Hospitals (Eur Heart J. 2015 Jul 17).
What makes this infections special, is the fact that the time to infection may takes months to years and that the micro-organism in question is easily missed by routine bacterial diagnostics.
The word is out, that other, difficult to diagnose micro-organisms e.g. Legionella are possibly causing post-operative infections, too. Thus, I believe that we can expect more cases with different pathogens in the near future.