There has been a lot of excitement about the prospects of whole genome sequencing (WGS) to support infection prevention and control in a really meaningful way over the past decade or two. But to me this potential seems largely unfulfilled. WGS remains largely the domain of reference and research laboratories, and has not transitioned effectively to support IPC daily decision making. A recent review highlights the potential of WGS to support IPC, and identifies some of the barriers to be overcome if WGS really is to be a big part of our future in IPC.
Continue readingWGS
Can you GES which carbapenemase caused this CPE outbreak?
An unusual and interesting outbreak of CPE was published recently in Clinical Infectious Diseases. Several key points: don’t rely solely on a PCR detecting the “Big 5” carbapenemases (NDM, KPC, OXA-48, IMP, VIM) – at some point you need to test for phenotypic carbapenemase activity; WGS can really help us in unravelling complex transmission routes; and covert plasmid propagation within and between species is a reality.
21 is the magic number (for defining CPE person-to-person transmission using WGS)
A fascinating study from a European research group has unravelled the molecular epidemiology of a large European collection of carbapenem-resistant Klebsiella pneumoniae clinical isolates. Most carbapenem resistance was due to an acquired carbapenemases, transmission clusters were common within and between hospitals, carbapenemase-producing isolates are more likely to spread in hospitals, and 21 SNPs is the magic number for defining CPE person-to-person transmission using WGS.
The importance of patient sharing between hospitals on MRSA transmission
In a remarkable quirk of academic publishing, two virtually identical studies by separate research groups in the UK (one in London, and one in Cambridge) published a week apart have come to the same conclusion: that we are missing a sizable portion of MRSA transmission by focussing solely on wards in a single hospital. A referral-network level view is required for an accurate picture of MRSA transmission. (You may have seen some press about the Cambridge article, e.g. on the BBC here.)
How much S. aureus is hospital acquired? Mk II
I posted a blog a couple of years ago (was it really that long!) on a fascinating study suggesting that only 1/5 of S. aureus in hospital patients is hospital-acquired. My key conclusion from that study was that the number of potential sources for S. aureus that the team investigated was inadequate to draw any firm conclusions (they didn’t include staff, surfaces, or visitors). I concluded that ‘the next frontier of transmission mapping must be a more comprehensive evaluation of other potential sources…’. The authors must have been reading, because this study from the same group was published recently in Lancet ID, which is a more comprehensive evaluation of other potential sources.
The carbapenemase is out there
A PNAS paper on the genomic diversity of carbapenemase producing bacteria in the US reports strong evidence of carbapenamase (an enzyme produced by bacteria that breaks down carabapenem antibiotics) activity but no sign of a carbapenemase gene. This provides a timely reminder that we are only really scratching the surface in our understanding of carbapenemases and how they work.
It’s transmission doc, but not as we know it
A groundbreaking study just published in PLOS Genetics provides new insight into the transmission dynamics of bacteria in the ICU setting using WGS. The ambitious authors performed WGS on virtually all bacterial isolates from ICUs in a US hospital for a year. The first surprise was that 12% of the bacteria considered clinically relevant were previously undescribed.
The next – and perhaps biggest – surprise was that whilst transmission of the usual suspect pathogens (MRSA, VRE etc) was rare, 9% of the other bacteria were shared by multiple patients, often with overlapping admissions (see the figure below). This suggests that there is a fair bit of transmission going on under the radar in the ICU setting.
Figure: Clonal lineages extending across multiple patients.
This study reminds me of one published in CID a few years ago showing that outbreaks of resistance probably occur regularly and usually undetected across multiple species.
So, is it time to start using WGS for all bacteria identified in the clinical laboratory? Not quite yet I don’t think: the analytical methods have not yet caught up with the sequencing technology. But this study is a glimpse of the future, no doubt about it.
Christmas 2014 Update
Now that you have digested your Christmas turkey, I thought that it would be a good time to send out an update. These articles have been posted since the last update:
- Who’s harbouring CRE? (Published 22nd December 2014)
- ECDC data shows progressive, depressing increase in antibiotic resistance in Europe (Published 16th December 2014)
- Is deliberately seeding hospital rooms with Bacillus spores a good idea? No, I don’t think so either! (Published 8th December 2014)
- Filling the gaps in the guidelines to control resistant Gram-negative bacteria (Published 2nd December 2014)
- Journal Roundup November 2014: Journal Roundup: Ebola (again), The rise (and rise) and fall of MRDOs & Infection Prevention 2014 (Published 28th November 2014)
- The inanimate environment doesn’t contribute to pathogen transmission in the operating room…OR does it? (Published 27th November 2014)
- Being bitten by antibiotic resistant CRAB hurts! (Acinetobacter that is.) (Published 25th November 2014)
- Reflections from HIS 2014, Part III: Education, communication, and antibiotic resistance (Published 21st November 2014)
- Reflections from HIS 2014, Part II: Dealing with the contaminated environment (Published 20th November 2014)
- Reflections from HIS 2014, Part I: Updates on C. difficile, norovirus and other HCAI pathogens (Published 19th November 2014)
- What’s trending in the infection prevention and control literature? (Published 16th November 2014)
- HIS Poster Round: Dealing with contaminated hands, surfaces, water and medical devices (Published 15th November 2014)
- Carbapenem-resistant Enterobacteriaceae (CRE): so what should an infection prevention and control team do now? (Published 11th November 2014)
- A postcard from Portugal: “Some days we don’t have any needles on the ICU” (Published 5th November 2014)
- Ebola: infection prevention and control considerations (Published 30th October 2014)
- ID Week 2014 as seen by an Infection Preventionist (Published 28th October 2014)
- Selective Digestive Decontamination (SDD) is dead; long live faecal microbiota transplantation (FMT) (Published 27th October 2014)
I’m in a rather reflective mood, so time to remind you of some of the key themes from 2014: Ebola, MERS-CoV, universal vs. targeted interventions, faecal microbiota transplantation (FMT), whole genome sequencing (WGS), carbapenem-resistant Enterobacteriaceae (CRE), and some interesting developments in environmental science. And what will we be still talking about come Christmas 2015? Let’s hope it won’t be Ebola, and I think that WGS will be a lab technique akin to a Vitek machine rather than subject matter for NEJM. But I think we still have ground to cover on whether to go for universal or targeted interventions, FMT, and improving our study designs in infection prevention and control. I can also foresee important studies on the comparative and cost-effectiveness of the various tools at our disposal.
And finally, before I sign off for 2014, a classic BMJ study on why Rudolf’s nose is red (it’s to do with the richly vascularised nasal microcirculation of the reindeer nose, apparently).
Image: Christmas #27.
Reflections from HIS 2014, Part I: Updates on C. difficile, norovirus and other HCAI pathogens
The 2014 Healthcare Infection Society (HIS) Conference was in Lyon, France, and combined with SFH2 (The French Society for Hospital Hygiene). Congratulations to all involved (especially Martin Kiernan and Prof Hilary Humphreys) for such a stimulating programme, and enjoyable conference. The abstracts from the oral presentations can be downloaded here, and the posters here. I plan to share some of my reflections on key conference themes over the next few days:
- Part I: Updates on C. difficile, norovirus and other HCAI pathogens
- Part II: Dealing with the contaminated environment
- Part III: Education, communication, and antibiotic resistance
- ‘What’s trending in the infection prevention and control literature?
HIS 2012 -> HIS 2014’ - ‘HIS Poster Round: Dealing with contaminated hands, surfaces, water and medical devices.’
Prof Wing-Hong Seto – Airborne transmission and precautions – facts and myths
Prof Seto’s energy and enthusiasm lit up the stage, just like a few years ago in Geneva for ICPIC. Prof Seto spent his lecture convincingly debunking the idea that airborne transmission of respiratory viruses is common, notwithstanding some data that, prima facie, suggests this. Only very few pathogens require obligate airborne transmission (e.g. TB); some have preferential airborne transmission (e.g. measles); and some have potential airborne transmission (respiratory viruses). There is some evidence that respiratory viruses such as influenza can be transmitted via the airborne route, but the most important route of transmission will depend on context. One important point is that studies demonstrating airborne “transmission” using PCR rather than viral culture as an endpoint, or using artificial aerosol generation should not be taken as definitive evidence of airborne transmission. Prof Seto’s view is that medical masks are sufficient to prevent the transmission of respiratory viruses, as demonstrated by his own work during SARS. Finally, we can forget the requirement for negative pressure isolation rooms: open doors and windows yields a whopping 45 air changes per hour!
Prof Mark Wilcox – Is Clostridium difficile infection (CDI) underestimated due to inappropriate testing algorithms?
Prof Wilcox began by reporting an unusual epidemic: “PCRitis”, which can cloud rather than clarify accurate diagnosis of CDI. Perhaps the most important point made by Prof Wilcox is that the ultimate “gold standard” for CDI should be clinical, and not laboratory based. Prof Wilcox spent most of his time reflecting on the recent multicentre European study of CDI underdiagnosis in Europe. There are some real shockers in here: the reported rate of CDI in Romania was 4 cases per 1000 patient days vs. closer to 100 per 1000 patient days when samples from the same patients were tested in the reference lab. This is no surprise in a sense because only 2/5 local laboratories were using optimal methods. However, even in the UK where around 80% of local labs are using optimal methods, around 2-fold more cases were identified in the reference vs. the local laboratory. Clearly, if we’re going to have a hope of controlling the spread of C. difficile in Europe, laboratory diagnosis needs to improve.
Norovirus
Norovirus is especially topical in the UK given the recent PHE announcement about unusually high rates of norovirus in the NHS. The prolific Dr Ben Lopman (CDC) began by explaining the ‘image problem’ that norovirus has in US hospitals, where it is considered an uncommon cause of gastroenteritis. In fact, a systematic review found that norovirus cases around 20% of acute gastroenteritis. However, I would say it’s just not possible to get an accurate assessment of how common norovirus is on a population level due to chronic under-reporting. When we had an outbreak of ”norovirus” in the Otter household, the last thing we felt like doing was submitting a specimen, and I suspect we are not alone in this! Although norovirus is usually mild and self-limiting, it is by no means benign: one Lopman study suggested that it is responsible for 20% of deaths due to gastroenteritis not caused by C. difficile in those ages >65. And then there’s the infection control challenges. Due to the exquisitely low infectious dose, 2g of stool from an infected individual is enough to infect the entire human population! Plus, it is shed in high titre, stable in the environment, and resistant to many disinfectants. Rather depressingly, it seems that effective interventions to control norovirus teeter around the cost-effectiveness threshold. More optimistically though, prospects for vaccines look promising.
Prof Marion Koopmans then described the huge diversity within the “norovirus” family, spanning more phylogenic space than many single species occupy. For chapter and verse on nomenclature, see Norovirus Net. It’s difficult to know what works to control norovirus due to dynamic outbreak settings combined with multiple interventions. One key aspect for control is understanding shedding profiles of infected, recovered and asymptomatic individuals. Whilst all can shed norovirus, much like Ebola, those who are symptomatic are by far the highest risk for transmission. Finally, our inability to culture norovirus in the lab has been an important barrier to understanding the virus; a recent study (in Science no less) suggests that a working lab model for culturing norovirus may be just around the corner.
Dr Lennie Derde – Rapid diagnostics to control spread of MDR bacteria at ICU
Given the turnaround times of conventional culture (24 hours to preliminary results – at best), rapid PCR-based diagnostics make sense in principle. But do they work in practice? There is some evidence that rapid diagnostics may work to reduce MRSA transmission, although other studies suggest that they don’t make a difference. In order to put rapid diagnostics to the test Dr Derde et al. ran the impressive MOSAR study. This study suggest that screening and isolation by conventional or rapid methods does not help to prevent the transmission of MDROs in the ICU, but I don’t think we should take that away from this study, not least due to the fact that many units were already doing screening and isolation during the baseline period!
New insights from whole geneome sequencing (WGS)
WGS is trendy and trending in the infection prevention and control sphere. Prof Derrick Crook gave an engaging overview of the impact that WGS has made. It’s analogous to the manual compilation and drawing of maps to GPS; you wouldn’t dream of drawing a map by hand now that GPS is available! Desktop 15 minute WGS technology will be a reality in a few years, and it will turn our little world upside down. The major limiting step, however, is that mathematics, computer science and computational biology are foreign to most of us. And we are foreign to most of them! But, these issues are worth solving because the WGS carrot is huge, offering to add new insight into our understanding of the epidemiology of pathogens associated with HCAI. For example, Prof Crook WGS study on C. difficile suggests that transmission from symptomatic cases is much less common than you’d expect. So if the C. difficile is not coming from symptomatic cases, where is it coming from? Contact with animals and neonates in the community are plausible sources However, I was surprised that Prof Crook didn’t mention the large burden of asymptomatic carriage of toxigenic C. difficile, which must be a substantial source for cross-transmission in hospitals.
WGS has yielded similar insight into the epidemiology of TB and MRSA, outlined by Drs Timothy Walker and Ewan Harrison, respectively. One challenging idea from Dr Harrison is how much of the “diversity cloud” that exists within an individual is transferred during a transmission event? Finally, WGS can turn a ‘plate of spaghetti’ of epidemiological links to a clear transmission map, as was the case during a CRE outbreak at NIH in the USA.
Look out for some more reflections from HIS posted over the next few days…
How much patient-to-patient spread of S. aureus occurs? Apparently, not much
I recently posted an article on surprising finds of a study suggesting that horizontal transmission of C. difficile from known symptomatic cases may be less common that we thought. A group of researchers from Oxford, Brighton and London in the UK applied similar methodology to Staphylococcus aureus transmission with similar findings: only a fifth of S. aureus acquisitions could be attributed to patient-to-patient transmission.
All patients admitted to a 16 bed ICU in Brighton were screened on admission and weekly to detect S. aureus colonization and acquisition. Each isolate was typed by spa and whole genome sequencing (WGS). The number of acquisitions that could be linked using conventional methods (spa typing combined with an analysis of overlapping stays) vs. WGS was evaluated.
Overall, 185 (16.7%) of 1109 admissions carried S. aureus; 59 carried MRSA (5.3%). 680 patients were on the unit for long enough to have a weekly screen and hence were eligible for assessing acquisition. Of these, 44 S. aureus (22 MRSA) acquisitions were detected in 41 patients. 35 of these acquisitions were in patients who were screen-negative on admission and 9 were acquisitions of different strains in patients who were already colonized on admission.
Only 14% (5/36) of the acquisitions available for typing were of the same spa type as another patient on the unit at the same time. All of these were MRSA. WGS discounted three of these apparent occurrences of patient-to-patient transmission, confirmed two and identified a further five (3 MRSA, 2 MSSA). So, in total, 7 / 36 (19%) of acquired isolates (5 MRSA and 2 MSSA) were linked to isolates from other patients (Figure).
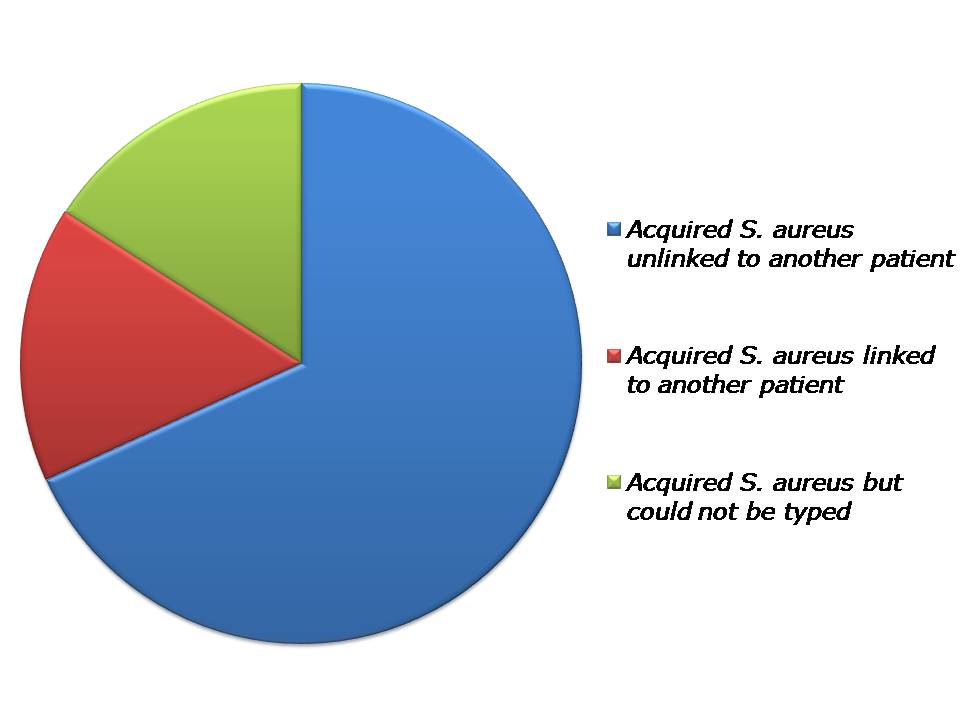 Figure: Source of S. aureus acquisitions identified through WGS.
Figure: Source of S. aureus acquisitions identified through WGS.
The paper raises some interesting questions:
- Principally, if almost 80% of patients did not acquire their S. aureus from other patients on the unit, where on earth did they acquire them from? In the C. difficile study, unrecognized importation of diverse strains from the community that would not be detected on admission since admission screening was not performed represents a plausible explanation for the surprisingly low incidence of horizontal transmission from known cases. This is not the case in this study, where all patients were screened on admission for S. aureus. So where were the diverse isolates acquired from? Staff carriers? Contaminated surfaces? S. aureus has the capacity to survive on surfaces for very long periods (more than 1 year), so an ancient environmental reservoir is possible. Furthermore, there was no ‘lead in’ period to the study, so it could be that S. aureus on the unit in the months before the study left an environmental reservoir that led to acquisition in some cases, which would not have been captured by this study. The fact that 4/7 patient-to-patient transmissions acquired the same strain of S. aureus without sharing time on the ICU together supports this, although could also be explained by staff carriers. It was a shame that 16% of the 329 S. aureus isolates (including 16% (7/44) of the acquired isolates) were not available for sequencing, which represents a substantial reservoir from which patient-to-patient transmission likely occurred in some cases.
- It was interesting that some 20% (9/44) acquisitions that were detected occurred in patients who were already colonized on admission; these would have been missed altogether if molecular typing was not performed. I wonder how much ‘silent’ acquisition of this type occurs?
- Assuming temporal relationship between strains assumes a constant mutation rate. The ‘speed of the mutation clock’ was assessed in this study through repeated sampling of the same patient. This exercise demonstrated minimal diversity, as was the case for C. difficile.
- WGS is rapidly becoming the gold standard for transmission mapping. In this study, the conventional approach of evaluating spa typing with overlapping stays lacked both sensitivity and specificity for identifying transmitted isolates.
In summary, the major finding is that only 20% of patients who acquired S. aureus appeared to acquire it through horizontal spread from other patients. The next frontier of transmission mapping must be a more comprehensive evaluation of other potential sources: contaminated surfaces, contaminated air, nurses, doctors, cleaners, tea-tray deliverers and the list goes on…




