As England moves away from confirmatory PCR testing following a positive lateral flow test in the absence of COVID-19 symptoms, it’s a good time to look at what these two different testing strategies can offer us. There’s an excellent short review in NEJM combined with a case study to help illustrate the impact of pre-test probability plays out. Both lateral flow testing and PCR testing have their place, and in some ways lateral flow testing is a better correlate for infectivity (as well as being cheaper and easier!).
Continue readingPCR
HIS webinar – Understanding hospital-onset and hospital-acquired COVID-19 infection
The second in the series of excellent Healthcare Infection Society (HIS) interactive audience-led webinars went out a few weeks ago. The theme for this was hospital-onset and hospital-acquired COVID-19 infections, and here’s the video.
Testing patients and staff without symptoms for SARS-CoV-2: beware false-positives (and negatives)
As we begin to look to the other side of the peak of COVID-19, this issue of more widespread testing of patients, staff (and indeed the general population) for infection with the SARS-CoV-2 virus whether or not they have symptoms is looming large. We need to think carefully about the risk of false positives when interpreting the meaning of a positive PCR test in a group of people with a low prevalence of SARS-CoV-2.
CPE carriage: a rare event in London?
My old CIDR team have just published a study in JAC reporting a very low rate of carriage of CPE in patient admitted to a hospital in central London (just 5 (0.1%) of 4006 patients). This was a lot lower than we expected! Despite the very low rate of carriage, overseas hospitalisation was a significant risk factor for CPE carriage, and supports that we should be screening patients with recent overseas hospitalisation for CPE carriage.
MERS joins the more-environmental-than-you-may-think club
I blogged about a review of the surprising ability of some respiratory viruses (especially SARS-CoV and Influenza virus) to survive on dry surfaces last year. In the review, I predicted that MERS-Cov would also share the same ability to survive on dry surfaces as SARS-CoV – so I was interested to see a recent article in CID demonstrating that MERS is indeed more environmental than you may think.
Appraising the options for detecting carbapenemase-producing organisms
Carbapenemase-producing organisms (including CPE) present important clinical challenges: the “triple threat” of high levels of antibiotic resistance, virulence, and potential for rapid spread (locally, regionally, nationally, and globally)! However, these organisms somewhat ironically also present challenges to detection in the clinical laboratory. You’d expect that since these organisms are so important clinically they’d be dead easy to detect in the clinical lab – but this isn’t the case.
A comprehensive review published in Clinical Microbiology Reviews provides an overview of the diagnostic approaches to detect carbapenemase producers in the clinical lab. Figures 6 and 7 of the review provide a useful overview of the two broad approaches you could take: culturing organisms on agar plates, or using nucleic acid amplification techniques (NAAT – most commonly PCR) directly from a rectal swab.
Molecular diagnostics for C. difficile infection: too much of a good thing?
A study in JAMA Internal Medicine suggests that we may be ‘overdiagnosising’ C. difficile in this era of molecular diagnostics. The researchers from California grouped the 1416 patients tested for C. difficile into three groups: Tox+/PCR+ (9%), Tox-/PCR+ (11%), and Tox-/PCR+ (79%) (see Figure). Perhaps unsurprisingly, compared with Tox+/PCR+ cases, Tox-/PCR+ cases had lower bacterial load, less prior antibiotic exposure, less faecal inflammation, a shorter duration of diarrhoea, were less likely to suffer complications, and were less likely to die within 30 days. Perhaps even more importantly, patients with Tox-/PCR+ were pretty much identical to patients with Tox-/PCR- specimens in all of these metrics. In short: these patients had C. difficile in their gut, but they did not have C. difficile infection. The key message here is that we should not be treating patients who are C. difficile “positive” by molecular tests only.
How big is C. difficile infection in the USA?
 The New England Journal of Medicine recently published an article evaluating the burden of CDI in the USA. The huge CDC-led initiative collected data from 10 geographically distinct regions, identifying more than 15,000 cases. Around two-thirds of cases were classified as healthcare-associated (although only 25% were hospital-onset). This means that, prima facie, a third of CDI cases were community-associated. I find this proportion difficult to believe: I strongly suspect that many of these cases would have had healthcare-associated risk factors if the team were able to look hard enough. For example, they used a fairly standard 12 week look-back period to evaluate previous hospitalisation, but how would the data look if they’d used 12 months? Also, it’s usually only possible to evaluate previous hospitalisation in a single healthcare system, but many patients commute between various healthcare systems. The authors acknowledge in the discussion that this designation of “community-acquired” may be inaccurate based on the finding from a previous study whether healthcare-associated risk factors were identified in most patients, but only be a detailed phone interview.
The New England Journal of Medicine recently published an article evaluating the burden of CDI in the USA. The huge CDC-led initiative collected data from 10 geographically distinct regions, identifying more than 15,000 cases. Around two-thirds of cases were classified as healthcare-associated (although only 25% were hospital-onset). This means that, prima facie, a third of CDI cases were community-associated. I find this proportion difficult to believe: I strongly suspect that many of these cases would have had healthcare-associated risk factors if the team were able to look hard enough. For example, they used a fairly standard 12 week look-back period to evaluate previous hospitalisation, but how would the data look if they’d used 12 months? Also, it’s usually only possible to evaluate previous hospitalisation in a single healthcare system, but many patients commute between various healthcare systems. The authors acknowledge in the discussion that this designation of “community-acquired” may be inaccurate based on the finding from a previous study whether healthcare-associated risk factors were identified in most patients, but only be a detailed phone interview.
Scaling up from the figures from the 10 regions, national estimates were around 500,000 cases and 29,000 deaths due to CDI per annum in the US. This estimate is approximately double previous estimates for the national CDI burden in the USA, probably reflecting the adoption of molecular methods for the detection of CDI. This scaling up included an interesting statistical adjustment to see how prevalence varied depending on how many sites use sensitive molecular methods to detect CDI.
A sub-study included the culture of C. difficile from 1625 patients. More than 15% of stool specimens from patients diagnosed as CDI failed to grow C. difficile, probably illustrating the limitations of culture methods more than anything else. NAP1 (027) represented around half of cases, and was significantly more common in healthcare-associated CDI. I think it’s fair to say that the initial fears that NAP1 was a super-strain have been allayed by the fact that it’s now so common and there hasn’t been a surge in CDI mortality.
Finally, around 21% of healthcare-associated cases suffered at least one recurrence. Thus, there is a real need to the roll out of the uber successful faecal microbiota transplantation for recurrent CDI. In fact, there should be around 70,000 faecal microbiota transplantations each year in the US right now (500,000 x 0.66 x 0.21); I suspect there are far fewer.
CRE diagnosis: current status
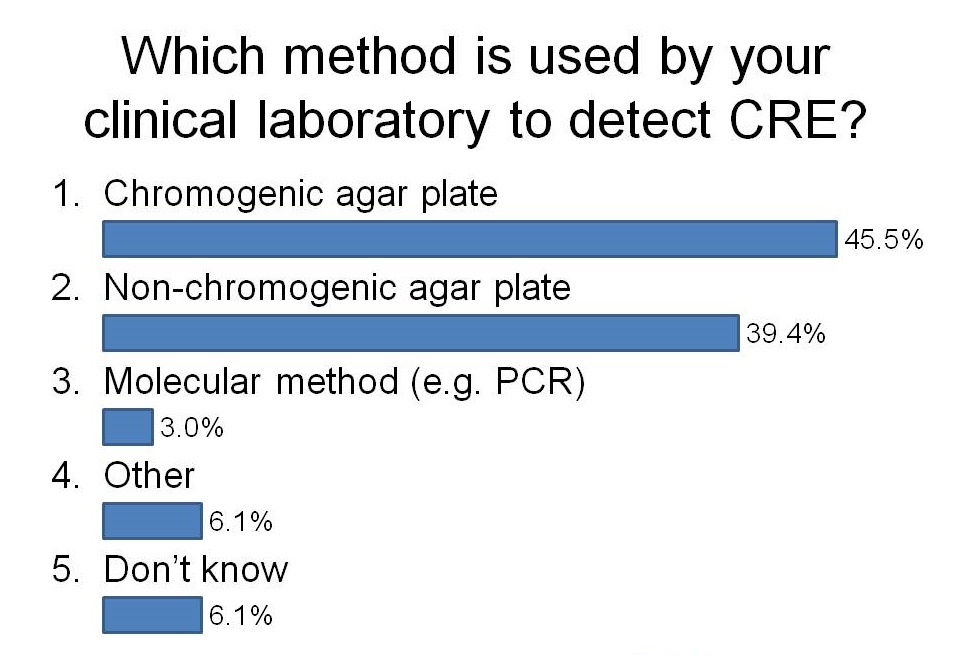
I had the opportunity to ask the audience how they were detecting CRE in their diagnostic clinical labs during a talk last week. It was an audience of around 50 laboratory and clinical folk, mainly from the UK but a few from continental Europe. And here’s what I found:
I was a little surprised that more labs have switched to using chromogeneic agar plates than use non-chorogeneic agar plates. In the case of our lab in London, we are currently using non-chromogenic media for clinical samples, but in the process of evaluating chromogenic media. Although the purchase costs of chromogenic media are higher, they are more sensitive and substantially reduce the amount of time required to confirm a negative or positive culture, so I suspect they actually work out cheaper when you factor in labour costs.
I was not surprised that so few labs are using PCR. The costs are considerably higher but turnaround time is faster and they are more sensitive. There are now a number of PCRs on the market for the detect of CRE direct from rectal swabs (e.g. Checkpoints and Cepheid). We are currently in the process of evaluating the Checkpoints assay and after sharing our preliminary data, this was the feeling in the room about using PCR to detect CRE:
I think I’ll leave it there for now…
Diagnosis of CRE: time to throw away those agar plates?
CRE are an emerging threat to healthcare systems worldwide. Most guidelines recommend screening and isolation of carriers. But relying on conventional agar-based culture presents a dual threat of poor sensitivity (depending on which method is used) and slow turnaround time, with a minimum overnight incubation before a presumptive positive result. PCR solves both of these problems – but at a cost. I gave a talk today at a BD seminar considering whether it’s time to switch to PCR diagnostics for the detection of CRE. You can download my slides here. (The title of the talk “Time to throw away those agar plates” was inspired by a talk by Dr Dan Diekema at a recent SHEA conference.)
There are a number of options for CRE screening, summarized in the flow chart below:
Flow chart: Overview of laboratory methods for the diagnosis of CRE. (To be precise, throughout the blog I really mean CPE most of the time, but I’m using CRE for consistency with other blogs…)