As England moves away from confirmatory PCR testing following a positive lateral flow test in the absence of COVID-19 symptoms, it’s a good time to look at what these two different testing strategies can offer us. There’s an excellent short review in NEJM combined with a case study to help illustrate the impact of pre-test probability plays out. Both lateral flow testing and PCR testing have their place, and in some ways lateral flow testing is a better correlate for infectivity (as well as being cheaper and easier!).
Continue readingdiagnostics
Dogs can be useful – Woof of concept obtained
I’m not a dog lover. Far from it in fact, however a new paper in the Journal of Hospital Infection caught my eye today. Yesterday I was sitting in the Longitude Prize Advisory  Committee meeting bemoaning the lack of ‘left field’ ideas coming forward. Harrison himself, winner of the original prize was such a person. He came at the problem of solving the longitude issue from a completely different direction when all of the respected science at the time was convinced that astrology was the answer. Problem: cloud, and not much of a silver lining. So we are looking for a new way to diagnose infection rapidly, distinguishing between those caused by viruses and bacteria in the hope of turning the increasing tide of resistance. So what does Fido (or Nimbus in this case) have to do with this?
Committee meeting bemoaning the lack of ‘left field’ ideas coming forward. Harrison himself, winner of the original prize was such a person. He came at the problem of solving the longitude issue from a completely different direction when all of the respected science at the time was convinced that astrology was the answer. Problem: cloud, and not much of a silver lining. So we are looking for a new way to diagnose infection rapidly, distinguishing between those caused by viruses and bacteria in the hope of turning the increasing tide of resistance. So what does Fido (or Nimbus in this case) have to do with this?
Review on AMR: final report
The final report from Jim O’Neill’s Review on AMR is published today. The report summarises the key findings of the reports published by The Review. The key interventions outlined in the report are:
- A global public awareness campaign
- Preventing the spread of infection
- Reducing unnecessary use of antibiotics, and controlling their environmental dissemination
- Improving surveillance of resistance and consumption
- Improving diagnostics
- Explore vaccines
- Improve remuneration for people working in ID (here here)
- Develop a global innovation fund for anti-infective drug development
- Incentivise anti-infective drug development
Not a great deal on infection prevention in the report – but this was covered in detail in a previous report. Some more excellent infographics, and an impressive Review. Well worth a read.
Review on AMR: progress to date
Like many others, I am keeping a close eye on the UK Government’s commissioned ‘Review on AMR’. The Review team have been tremendously productive over the last few years, already releasing detailed reports on:
CRE diagnosis: current status
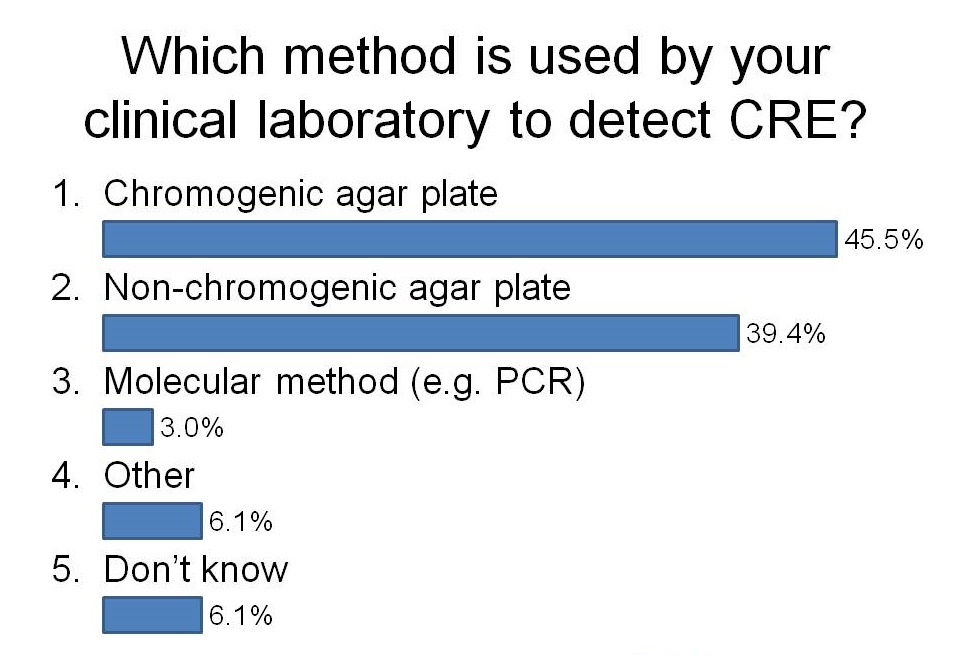
I had the opportunity to ask the audience how they were detecting CRE in their diagnostic clinical labs during a talk last week. It was an audience of around 50 laboratory and clinical folk, mainly from the UK but a few from continental Europe. And here’s what I found:
I was a little surprised that more labs have switched to using chromogeneic agar plates than use non-chorogeneic agar plates. In the case of our lab in London, we are currently using non-chromogenic media for clinical samples, but in the process of evaluating chromogenic media. Although the purchase costs of chromogenic media are higher, they are more sensitive and substantially reduce the amount of time required to confirm a negative or positive culture, so I suspect they actually work out cheaper when you factor in labour costs.
I was not surprised that so few labs are using PCR. The costs are considerably higher but turnaround time is faster and they are more sensitive. There are now a number of PCRs on the market for the detect of CRE direct from rectal swabs (e.g. Checkpoints and Cepheid). We are currently in the process of evaluating the Checkpoints assay and after sharing our preliminary data, this was the feeling in the room about using PCR to detect CRE:
I think I’ll leave it there for now…
CRE and friends: Q&A
 I gave the first in a three part webinar series for 3M last night, and you can download the slides here. Also, you can access the recording here (although you will need to register to do so).
I gave the first in a three part webinar series for 3M last night, and you can download the slides here. Also, you can access the recording here (although you will need to register to do so).
- Aug 19: CRE and friends: what’s the problem and how to detect them?
- Sept 16: Not all resistant Gram-negative bacteria are created equal: Enterobacteriaceae vs. non-fermenters
- Oct 7: Filling the gaps in the guidelines to control resistant Gram-negative bacteria
The webinar was attended by >200 participants from across the US. I tried to outline the three pronged threat of multidrug-resistant Gram-negative rods (especially CRE) in terms of high levels of antibiotic resistance, stark mortality (for invasive disease) and the potential for rapid spread (including the prospect of establishing a community reservoir). Then, I gave an overview of the US and European picture in terms of CRE prevalence. Finally, I discussed the diagnostic challenges and options.
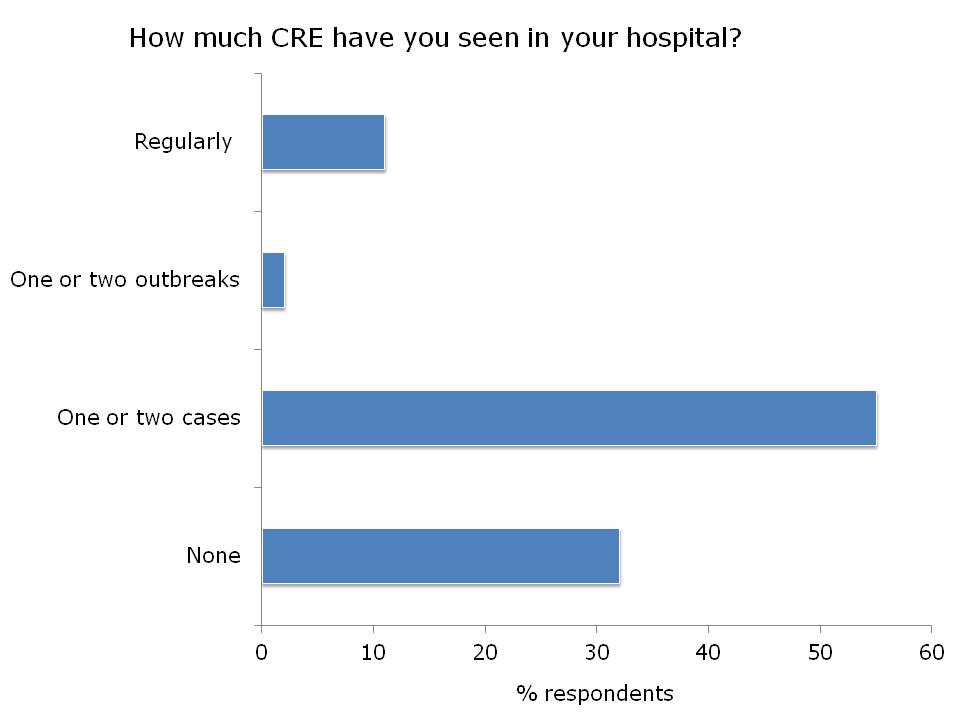
The most interesting part for me was the response to the questions that I threw out to the audience (see Figure below).
Figure: response to the questions from the 200 or so participants.
I was somewhat saddened but not especially surprised that the difference between CRE and CPE was not clear in the minds of most participants. I appreciate that this may be in part due to the fact that ‘CPE’ seems to be used more commonly in Europe than in the US. But this is an international problem, so we need to get our terminology straight in our globalised world.
It was encouraging to hear that most US hospitals have had no CRE, or only one or two cases. However, 11% of the participants see CRE regularly, with cases unconnected to outbreaks. This is a concern, and suggests that CRE has become established in these areas. Indeed, a recent study from 25 Southeastern US community hospitals reports a 5-fold increase in the prevalence of CRE since 2008, suggesting that CRE is becoming established in some parts of the US.
Most participants didn’t know which method was used by their clinical laboratory to detect CRE. I’m not sure whether or not this is a problem. You’d hope that laboratorians to know that they’re doing!
Q&A
The webinar included time for a Q&A from the audience, which covered the following:
- “How long to resistant Gram-negatives survive on surfaces?” This depends on which Gram-negative you’re talking about. Non-fermenters, especially Acinetobacter baumannnii, have remarkable survival properties measured in months and years. Enterobacteriaceae have a somewhat lower capacity to survive on dry surfaces, but it can still be measured in weeks and months, rather than hours and days.
- “How important is the environment in the transmission of resistant Gram-negatives?” Again, this depends on which Gram-negative you’re talking about. For A. baumannii the answer is probably “very important” whereas for the Enterobacteriaceae the answer is more like “quite important”.
- “What would you recommend for terminal disinfection following a case of CRE?” I would recommend the hospitals usual “deep clean” using either a bleach or hydrogen peroxide disinfectant, and consideration of using an automated room disinfection system. I would not be happy with a detergent or QAC clean; we can’t afford to leave an environmental reservoir that could put the next patient at risk.
- “Are antibiotic-resistant Gram-negative bacteria also likely to be resistant to disinfectants” There’s been a lot of discussion on this issue, but the short answer is no. I’d expect an antibiotic-resistant Enterobacteriaceae isolate to be as susceptible to disinfectants as a corresponding antibiotic-susceptible isolate.
- “Should patients with CRE be left to the end of surgical lists, and are is special instrument reprocessing required?” There is no need to implement special instrument reprocessing – follow your usual procedures here. Should CRE patients be left to the end of surgical lists? It would be prudent if possible, but don’t lose sleep over it.
- “Are any special decontamination measures necessary for endoscopes?” A number of outbreaks of CRE have been reported associated with endoscopy. However, usual endoscope reprocessing methods should be sufficient to deal with CRE, provided they are done correctly!
- “How do you lessen your chances of acquiring CRE?” Healthy individuals lack the risk factors for CRE infection (although CRE can occasionally cause infections in the community). Thus, the personal protective equipment (PPE) specified for contact precautions (gloves and gowns) combined with rigorous hand hygiene are sufficient to protect healthcare workers.
- “Are toilet seats in India safe?” What a question! I guess we’re talking about an organism with gastrointestinal carriage, so it’s probably that contamination of the toilet seat will occur. It may be prudent to clean or disinfect toilet seats in India before using them. Either that, or squat!
- “Can you expand on isolation protocols?” Firstly, ensure that patients infected or colonized with CRE are assigned a single room (not so relevant in the US, but important in healthcare elsewhere). Then, make sure you have appropriate policy and supply of PPE (principally gloves and gowns). Consider implementing ‘enhanced precautions’, including a restriction of mobile devices. Finally, consider cohorting patients and staff to the extent possible. You can read more about NIH’s approach to isolation here.
- “Can patients who are colonized with CRE be deisolated?” This is a tricky one, which is basically an evidence free zone and hence an area of controversy. Longitudinal studies show that carriage of CRE can persist for months or even years, so it makes sense to continue isolation for the duration of a hospitalization and not bother with repeated swabbing. At the time of readmission, it makes sense to take a swab to see whether colonization continues. If not, then it may be rational to deisolate them – perhaps after a confirmatory swab. I wish I could be more decisive here, but the evidence is scant.
Do please let me know if you have anything to add to this Q&A!
Journal Roundup July 2014
The July Journal Roundup is now available at the Journal of Hospital Infection website.
Topics this month include:
- The Longitude Prize.
- Randomized controlled trials of two novel glycopeptide antibiotics.
- Developments in antimicrobial therapy, including several new approaches to augmenting the activity of existing agents.
- Using chemicals for the prevention and treatment of skin wounds.
- Updates on antibiotic cycling (which does seem to work afterall).
- Commentary on ‘Mass Gatherings Medicine’.
- Further updates to the SHEA Compendium.
- Consideration of ‘horizontal’ (universal) vs. ‘vertical’ (targeted) strategies.
- A randomized controlled trial on the effectiveness of issuing mobile phone reminders for HIV appointments.
- A video surveillance study reporting a truly shocking level of hand hygiene compliance among anesthesiologists: 2.9%!
- Reviews on colistin, rapid nucleic acid based diagnostics, and the hithertofore unrecognised importance of free living amoebae in some healthcare-associated infection.
- And finally…what makes Twitter light up with antibiotic chat more than anything else?
Enjoy, and let me know if you have any questions or comments.
The pitfalls of PCR for detecting pathogens on surfaces
PCR has proven an invaluable tool for the rapid diagnosis of a range of pathogens, including MRSA and C. difficile. Several studies have evaluated the potential use of PCR for the detection of pathogens on surfaces and have identified some issues that, frankly, seem pretty terminal for this application using currently available commercial PCR kits.
A study from Cleveland evaluated the use of a commercial RT-PCR test for detecting C. difficile on hospital surfaces. Three composite sites were sampled in 22 patient rooms, 41% of which housed a patient with CDI with the remaining 59% sampled after terminal cleaning and disinfection. Two swabs and a gauze were collected from each site; one swab was cultured directly onto selective agar and the other was tested using PCR. The gauze was cultured using broth enrichment. C. difficile that grew on the selective agar were tested for toxin production and only toxigenic C. difficile were included.
Overall, 23 (35%) of the 66 sites grew toxigenic C. difficile and only 4 of these were detected using the standard RT-PCR assay (sensitivity 17%, specificity 100%). The sensitivity of RT-PCR in rooms that had been cleaned and disinfected was even worse (10%). Increasing the CT threshold of the assay (making it less stringent) improved the overall sensitivity to 52% and did not affect the specificity.
The study has several important limitations. The RT-PCR assay detected only the Toxin B gene, whereas the toxigenic culture methodology would detect both Toxin A and B producers. More importantly, there was a crucial difference in sampling methodology: the gauzes used for broth enrichment culture had a 50% higher positivity rate than the swabs (in line with other findings), but only swabs were tested by both PCR and culture. Thus, if the gauzes are a more effective sampling device, this would make the RT-PCR methodology seems worse than it is. I would have liked to have seen the sensitivity of the RT-PCR assay for detecting C. difficile cultured from the swabs only, but I could not derive this from the data in the paper.
An older study from New Haven, Connecticut provides a contrasting view of the use of PCR to detect pathogens from surfaces. Here, 10 standardized sites were sampled in the rooms of 10 patients infected or colonized with MRSA, and 5 rooms of patients not known to be infected or colonized with MRSA. Swabs were directly plated onto selective agar for MRSA, then DNA was extracted from the swabs before a broth enrichment procedure using the same swabs. In this study, 40 (27%) of the 150 surfaces were positive by culture, but 90 (60%) were positive by PCR (sensitivity 93%, specificity 51%).
Figure 1. Contrasting sensitivity and specificity when using PCR to detect C. difficile and MRSA on hospital surfaces.
It seems then that the sensitivity of PCR is too low for the environmental detection of C. difficile but the specificity is too low MRSA (figure 1). How could this be? Assuming that this is not due to experimental differences between the studies, it could be that the standard extraction procedure used for the C. difficile assay was not robust enough to liberate DNA from the mature environmental spores, resulting in low sensitivity. Conversely, the PCR assay was detecting DNA from dead MRSA on surfaces, resulting in low specificity.
So, in summary, the MRSA assay was too sensitive and the C. difficile assay was not sensitive enough! While the use of these “off the shelf” commercial assays doesn’t seem to be useful for detecting pathogens on surfaces, there may be hope for a PCR assay tailored specifically for an environmental application.
Article citations: