One of the faces of the global antibiotic resistance crisis is Escherichia coli ST131, frequently portrayed as a pandemic clone, combining hypervirulence, ciprofloxacin resistance and ESBL production. A recent study in Genome Research, a journal you may not read every month, though, sheds a whole new light on this “superbug”.
British and a Finnish researcher investigated 1,509 E. coli bacteremia isolates from the UK, collected in a non-selective manner, and spanning a time period of 11 years (2001-2011). Everything inside the bacterium that could be sequenced was sequenced. With that information E. coli were given their “international” names and their prevalence was followed in time. The most prominent types were ST173 and ST131. ST173 was already there in 2001 and managed to stay, i.e. it’s proportional contribution to all E. coli infections remained stable.
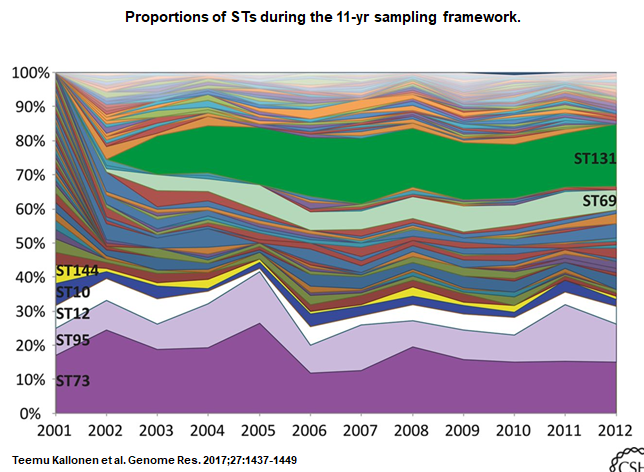
ST131 entered the arena in 2002, took its position in 2 years and then remained stable over time. Both types were in equilibrium. Now the funny finding is that most ST131 isolates were, as expected, resistant to Cipro and cephalosporins, whereas ST173 was not. So, it maintained its position, as ST131 did, without resistance. According to the authors it can, therefore, not be concluded that the emergence of ST131 was driven by antibiotic resistance traits.
Another interesting finding was that of all 3,511 virulence genes in the whole collection only 1 appeared specific to ST131. And apparently, ST131 became ST131 around 1874 (the year that Aston Villa F.C. was founded by members of Villa Cross Wesleyan Chapel cricket team in Handsworth, Birmingham, says Wikipedia). And the clade of ST131 characterized by Cipro resistance diverged in 1982 (the year Paulo Rossi defeated the most beautiful (non-Dutch) national football team ever).
So, if not hypervirulence and multiple resistance, what then explains ST131’s existence? The logical explanation – of the authors – is simple and called “negative frequency-dependent selection”. In laymen terms (my interpretation): A new kid on the block is successful as long as it is seen as a new kid, but its advantage is lost when becoming part of the establishment. Which fits with the scenario that E. coli bloodstream infections represent a spill-over of what is in the people’s gut, and the gut population is shaped in the community, where there is low antibiotic selective pressure, and where antibiotic resistance is not advantageous. It’s the opposite of MRSA, where the population structure of isolates causing infections is created in hospitalized patients, where drug resistance creates a strong selective advantage and pandemic clones have emerged.
Fantastic work. Although it may not be the last word on the faith of ST131 (and mankind), it tells us to be careful when calling a bug a superbug. If you have difficulties in believing genetic data, the same conclusion was reached after a systematic review and meta-analysis published in BMJ Open, see.
Discover more from Reflections on Infection Prevention and Control
Subscribe to get the latest posts sent to your email.